Sequence alignment is the procedure of comparing two (pairwise alignment) or more multiple sequences by searching for a series of individual characters or patterns that are in the same order in the sequences.
Global Sequence Alignment vs Local Sequence Alignment
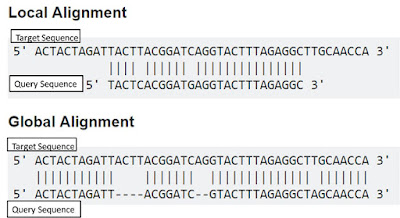
Global Sequence Alignment
|
Local Sequence Alignment
|
In global alignment, an attempt is made to align the entire sequence (end to end alignment)
|
Finds local regions with the highest level of similarity between the two sequences.
|
A global alignment contains all letters from both the query and target sequences
|
A local alignment aligns a substring of the query sequence to a substring of the target sequence.
|
If two sequences have approximately the same length and are quite similar, they are suitable for global alignment.
|
Any two sequences can be locally aligned as local alignment finds stretches of sequences with high level of matches without considering the alignment of rest of the sequence regions.
|
Suitable for aligning two closely related sequences.
|
Suitable for aligning more divergent sequences or distantly related sequences.
|
Global alignments are usually done for comparing homologous genes like comparing two genes with same function (in human vs. mouse) or comparing two proteins with similar function.
|
Used for finding out conserved patterns in DNA sequences or conserved domains or motifs in two proteins.
|
A general global alignment technique is the Needleman–Wunsch algorithm.
|
A general local alignment method is Smith–Waterman algorithm.
|
Examples of Global alignment tools:
|
Examples of Local alignment tools:
|
This is summarized video on the topic
Enroll Now - Understand
the Basics of rDNA Technology in 2 hours
References:
- https://www.ebi.ac.uk/Tools/psa/
- https://www.expasy.org/genomics/sequence_alignment
- https://blast.ncbi.nlm.nih.gov/Blast.cgi
Good One
ReplyDeleteThank you so much
ReplyDeletePost a Comment
We Love to hear from U :) Leave us a Comment to improve this site
Thanks for Visiting.....